At EMNLP 2024, the Best Paper Award was given to “Pretraining Data Detection for Large Language Models: A Divergence-based Calibration Method”
However, upon closer examination, we identified significant shortcomings in both the experimental design and evaluation methodology. The proposed dataset introduces a temporal shift between the distribution of member and non-member data, which can lead to overestimation of the performance of an MIA that may end up distinguishing samples based on the temporal range, and not actual membership.
In this post, we critically analyze this shift, and the broader implications of MIA evaluations for models in the wild.
What is Membership Inference?
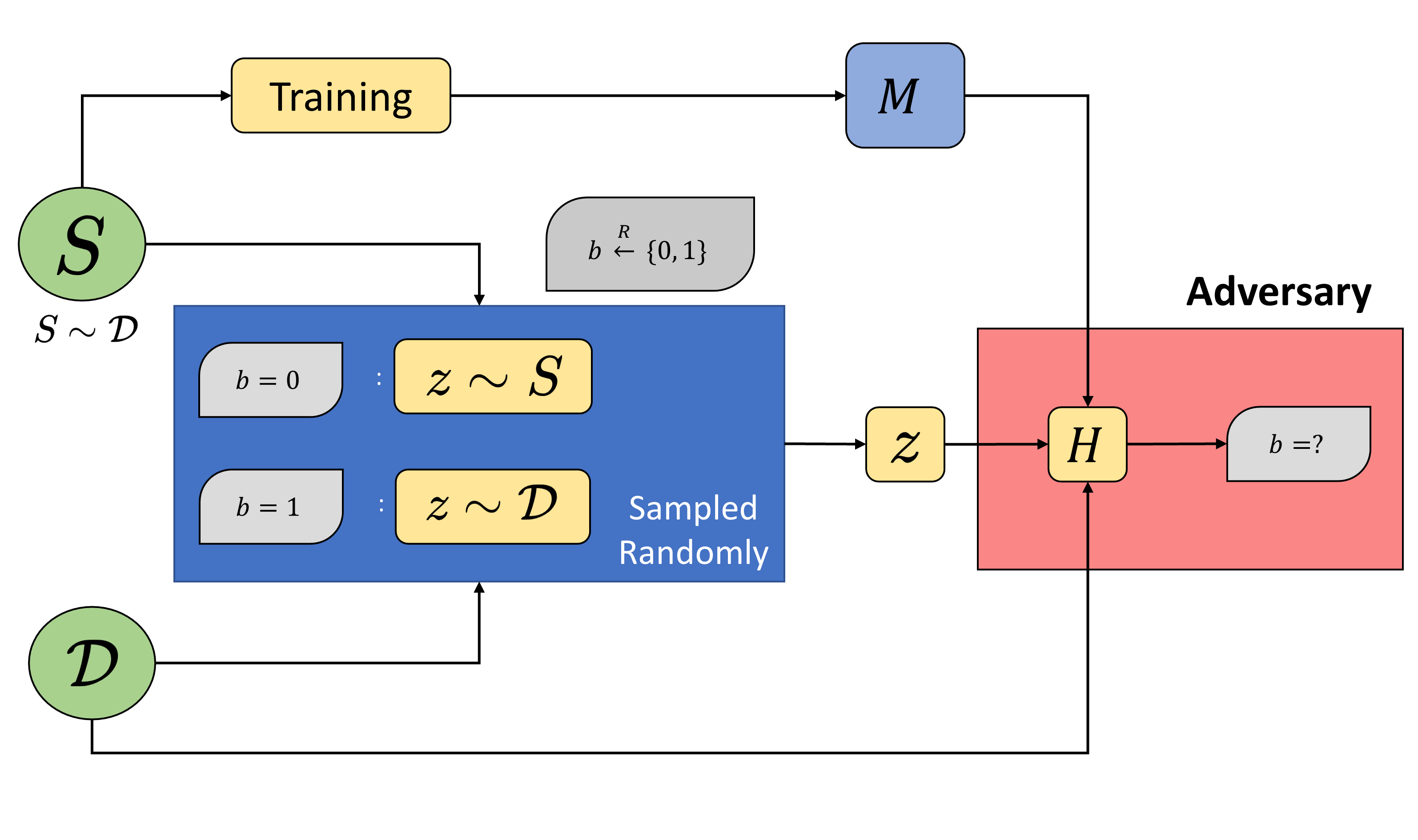
Membership Inference Attacks (MIAs) are a useful tool in assessing memorization of training data by a model trained on it. Given a model \(D\) samples from some underlying distribution \(\mathcal{D}\) and a model \(M\) trained on \(D\), membership inference
Was some given record \(x\) part of the training dataset \(D\), or just the overall distribution \(\mathcal{D}\)?
The underlying distribution \(\mathcal{D}\) is assumed to be large enough to the point where the above test can be reframed as inferring whether \(x \in D\) (via access to \(M\)) or not. In practice, the adversary/auditor starts with some non-member data (data that they know was not part of the training data \(D\), but belongs to the same underlying distribution \(\mathcal{D}\)) and on the basis of some scoring function, generates a distribution of scores for these non-members. A sweep over these values can then yield “thresholds” corresponding to certain false-positive rates (FPRs), which can then be used to evaluate the true-positive rate (TPR) of the approach under consideration.
It is important to note here that these non-members should be from the same underlying distribution. To better understand why this is important, think of a model trained for the binary classification task of distinguishing images of squirrels and groundhogs



A model will have higher loss on images of llamas, and understandably so since these are images the model did not see at all during training. Using their images would give a clear member/non-member distinction, but would also probably classify any squirrel/groundhog image as a member, even if it wasn’t. As an experimental setup, this is easily enforced when working with standard machine learning models and datasets such as CIFAR-10 and ImageNet, where well-established train/test splits from the same underlying distribution exist.
What’s Special about LLMs?
Because these models are trained on a large scale of data (and in many cases, exact training data is unknown), it is hard to collect data to use as “non-members” which has not been used in the model training and is from the same underlying distribution. Early works on membership inference for LLMs resorted to using data generated after a model’s training cutoff
Method Overview
The proposed method tries to fix a known issue with MIAs: models often fail to properly separate member and non-member samples. To address this, the authors use an auxiliary data-source to compute token-level frequencies, which are then used to recalibrate token-wise model logits. This normalization aims to adjust token-level model probabilities based on their natural frequency or rarity, aligning with membership inference practices such as reference model calibration
They also introduce PatentMIA, a benchmark that uses temporally shifted patents as data. The idea is to test whether the model can identify if a patent document was part of its training data or not. While this approach sounds interesting, our experiments suggest that the reported results are influenced by limitations in the benchmark design.
Experimental Evaluation
We ran two key experiments to test the paper’s claims: one for true positives and another for false positives.
True Positive Rate Experiment
This experiment checks if the method can correctly distinguish member data from non-member data when both are drawn from the same distribution. We used train and validation splits from The Pile dataset, which ensures there are no temporal or distributional differences between the two sets. Below we report results for the Wikipedia split.
| Model | AUC | TPR@5%FPR |
|---|---|---|
| Pythia-6.9B | 0.542 | 0.071 |
| GPT-Neo-125M | 0.492 | 0.054 |
| GPT-NeoX-20B | 0.600 | 0.103 |
Result:
The method performs only slightly better than the LOSS attack, and remains comparable to most standalone membership inference attacks. For reference, AUC with the baseline LOSS and zlib
Reported improvements in the paper (Table 2
False Positive Rate Experiment
Next, we check how often the method falsely identifies data as “member” when it has in fact not be used in the model’s training. To do this, we use the WikiMIA
Result:
Below we report results for the Wikipedia split. Note that in this setting, a score closer to 0.5 is better since both splits are non-members.
| Model | AUC for DC-PDD | AUC for LOSS |
|---|---|---|
| Pythia-6.9B | 0.667 | 0.636 |
| GPT-Neo-125M | 0.689 | 0.671 |
| GPT-Neox-20b | 0.637 | 0.656 |
The method flags a high number of false positives. It frequently identifies non-member data as part of the training set, suggesting that the attack was was reliant on temporal or distribution artifacts rather than truly detecting membership.
The Problem with Temporally Shifted Benchmarks
The introduction of PatentMIA highlights a broader problem with MIA research: benchmarks that rely on temporal shifts
Why These Benchmarks Are Misleading
The issues with temporally shifted benchmarks are not new. Several prior works have already established the dangers of using such benchmarks:
- Spurious Patterns: Temporal shifts introduce artifacts that are easily exploitable by attack models. As noted by Duan et al.
, temporal markers (e.g., “COVID-19” or recent events) allow models to cheat by detecting new concepts rather than true membership. - Misleading Evaluations: Maini et al.
show how temporal shifts can inflate the perceived success of MIAs, even when no meaningful membership inference occurs. - Blind Baselines Work Better: Das et al.
demonstrate that blind baselines often outperform sophisticated MIAs on temporally shifted datasets, highlighting how these benchmarks fail to test real inference ability.
Despite these well-established issues, the EMNLP Best Paper continues to rely on temporally shifted data like PatentMIA for its evaluations. This undermines the robustness of its claims and contributes little to advancing membership inference research.
Machine Learning Awards: A Problem of Incentives
This situation raises important questions about the role of awards in machine learning research.
- Do Awards Encourage Rushed Work? Highlighting work with known flaws, like relying on misleading benchmarks, can discourage researchers from investing time in more rigorous evaluations.
- Harming the Field: Awards that celebrate flawed work set a bad precedent and can mislead the community into thinking these methods are the gold standard.
- Losing Credibility: Over time, the reputation of awards themselves suffers, as researchers may start viewing them as less meaningful.
This is a growing problem in machine learning research, where not only acceptance but even awards are constantly under scrutiny for their soundness, let alone their contribution. If awards are to truly highlight excellence, they must emphasize thoroughness, reproducibility, and robustness over surface-level novelty.
Conclusion
The EMNLP 2024 Best Paper sought to address a pressing challenge in membership inference but falls short under careful scrutiny. The proposed method fails both in distinguishing members and non-members under rigorous conditions and in avoiding false positives when the data is untrained. Furthermore, its reliance on PatentMIA exemplifies a larger issue with using temporally shifted benchmarks to claim progress.
For the field to advance meaningfully, greater emphasis must be placed on rigorous evaluation practices. Awards should reflect this by rewarding work with robust and thorough evaluations, rather than methods that (knowingly or otherwise) exploit well-known flaws in evaluation practices. Only then can we ensure that the field moves forward in a meaningful way.
Acknowledgements
We would like to thank Zack Lipton and Zico Kolter for their helpful feedback on the draft and for referring us to Nicholas’s